# Introduction
The Niche Model Hegemony: Establish how the Niche Model (and its topological derivatives like the Cascade model) has historically served as the “structural prior” or template for most network theory.
The Philosophy Gap: Contrast structural/topological models (which prioritise species-agnostic link distributions) with “realised/dynamic” models like ATN and ADBM (which prioritise biological mechanisms like bioenergetics and optimal foraging).
The Research Question: Do these different philosophical starting points (mechanical vs. statistical) lead to the same conclusions about community stability, or does the choice of “template” fundamentally bias our understanding of extinction dynamics?.
1 Methods
1.1 Model Selection and Implementation
Describe the seven model classes - Structural: Niche, Cascade, MaxEnt (Topological templates) vs realised/dynamic: ATN (mechanical/allometric), ADBM (energetic/optimal foraging), and Latent Trait (statistical/allometric)
| Model | Type | Main Inputs | Key Assumptions |
|---|---|---|---|
| Niche model (Williams and Martinez 2000) | Structural | Species niche values | Trophic interactions structured by a 1D feeding niche; species consume all prey within a contiguous range; allows cannibalism. |
| Cascade model (Cohen et al. 1990) | Structural | Species rank | Consumers feed only on species with lower ranks; strictly hierarchical; no loops or omnivory. |
| MaxEnt model (Banville et al. 2023) | Structural | Species richness, connectance | Links assigned to maximize entropy; captures global network properties without species-specific mechanisms. |
| Allometric Diet Breadth Model (ADBM) (Petchey et al. 2008) | Realised / Dynamic | Body mass, prey energy content | Consumers maximize energy intake; diet breadth determined by profitability and handling time; allometric scaling governs parameters. |
| Allometric Trophic Network (ATN) (Brose et al. 2006) | Realised / Dynamic | Body mass | Interactions constrained by body-mass ratios; mechanical size limits structure networks; probability of interaction follows Ricker function. |
| Latent trait model (Rohr et al. 2010) | Realised / Dynamic | Species traits (e.g., body mass) | Interactions inferred statistically based on hidden trait correlations; can incorporate probabilistic constraints on links. |
Simulation Framework: Explain how an identical species pool was used across all models to isolate the model effect from the data effect. Something about how we constrained networks and also species class assignments if and when valid in constraining the model vs needed as input for the BEFW
Dynamic Perturbation: Detail the use of a bioenergetic food web model to simulate species loss and cascading extinctions.
Metrics for Comparison: Define the macro-scale (connectance), meso-scale (motifs), and micro-scale (generality/vulnerability) properties used to evaluate structural stories. As well as some of the outputs from the dynamic models - persistence/stability
2 Results
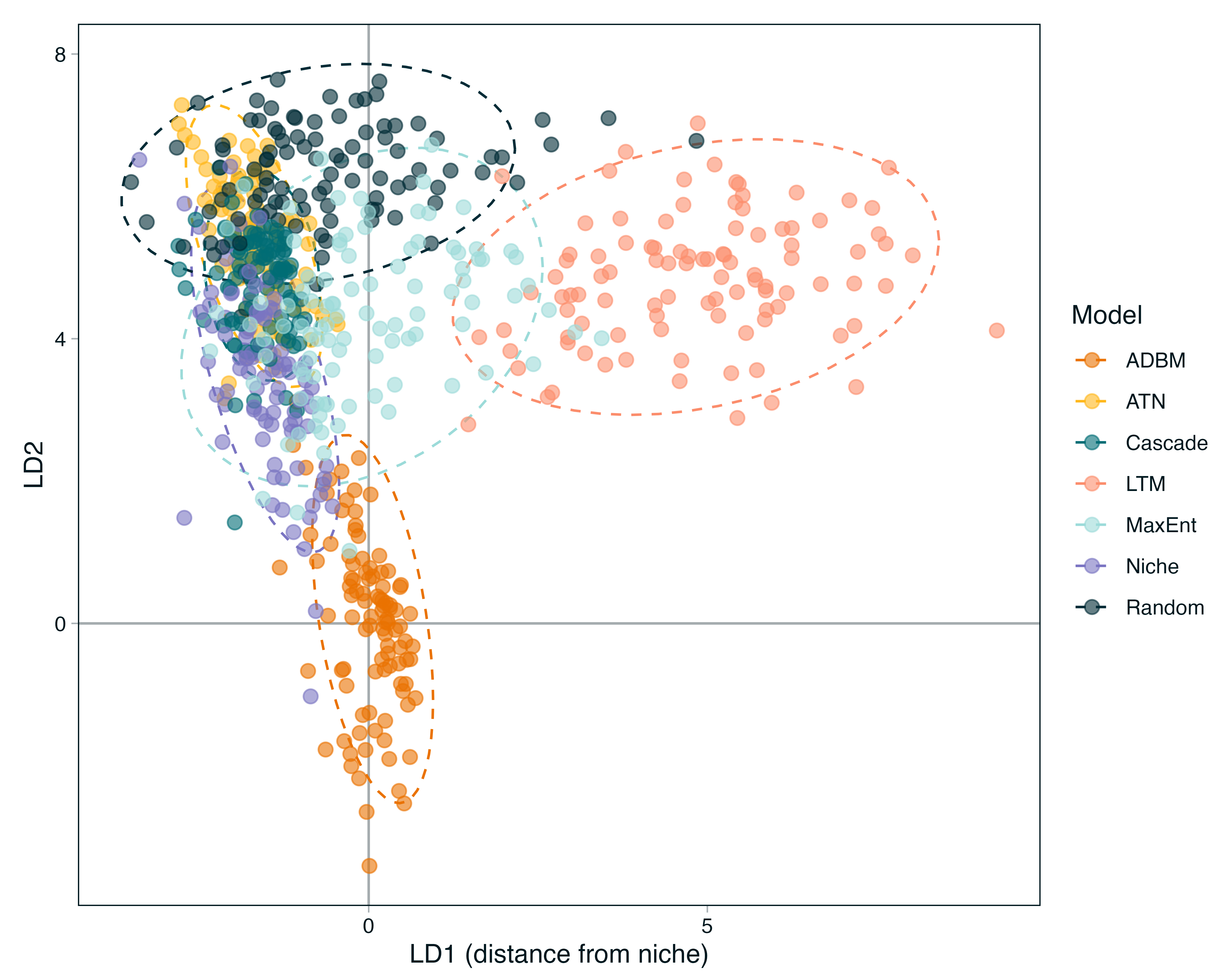
Structural Signatures: Report how much variance is explained by the model choice.
The Niche Model Template: Identify if the structural models occupy a central ‘mean space’ or if the dynamic models diverge into non-overlapping regions.
Divergent Stories in Dynamics: Show how models with similar global structures (like connectance) can lead to wildly different pathways of change/collapse/straight lines.
3 Discussion
Reconstruction is Not Neutral: Discuss how models act as structural priors that condition ecological inference. Well maybe - paleo webs seems to suggest this
Template Dependence: Address the core question: Is everything a derivative of the niche model? If dynamic models show unique cascade patterns not captured by the niche model, argue that the niche template may be insufficient for dynamic questions.
Scale-Dependent Robustness: Reflect on why species-level patterns might look similar across models while the fine-grained story of dynamic change is highly model-contingent.
Implications for Theory: Suggest that researchers must align their reconstruction framework with their inferential goals rather than defaulting to a one-size-fits-all niche template.