1 Introduction
Understanding how biodiversity is organised across space and time is a central goal of macroecology and biogeography. While early efforts focused primarily on species richness and composition, there is growing recognition that ecological communities are structured not only by which species occur, but by how they interact (Thuiller et al. 2024). Interaction networks are increasingly treated as macroecological state variables where they are used to compare community organisation across environmental gradients, to quantify β-diversity in interaction structure, to evaluate stability–complexity relationships, and to infer vulnerability under global change (Tylianakis and Morris 2017; Poisot et al. 2015; Gravel et al. 2019; Trøjelsgaard and Olesen 2016).
As a result, ecological networks now play a central role in comparative analyses spanning latitudinal gradients, disturbance regimes, and deep-time environmental transitions (Hao et al. 2025; Michalska-Smith and Allesina 2019; Dunhill et al. 2024; Roopnarine 2006; Poisot and Gravel 2014). Implicit in this expansion is the critical assumption that network properties estimated across systems are structurally comparable, and that differences among them reflect ecological signal rather than methodological artefact (Jordano 2016; Fründ et al. 2016).
However, most ecological networks are not fully observed as interaction data are incomplete and sampling is uneven across historical and biogeographic contexts, across both present day and deep-time (Sandra et al. 2025; Poisot et al. 2021). Interactions must often be inferred indirectly from traits, phylogeny, body size, co-occurrence, or theoretical constraints (Strydom et al. 2021; Morales-Castilla et al. 2015). Network construction therefore constitutes a model-based inference step rather than a purely descriptive exercise. Different reconstruction frameworks encode distinct ecological assumptions about how interactions arise - whether as biologically feasible combinations of traits, energetically optimised realised diets, or topological structures constrained by macroecological regularities. These assumptions act as structural priors over network architecture (Strydom et al. 2026; Gauzens et al. 2025; Petchey et al. 2011; Guimarães 2020). If alternative reconstruction approaches systematically generate different trophic configurations, then comparative analyses risk conflating ecological differences among communities with artefacts introduced by reconstruction choice. The reliability of macroecological inference therefore depends not only on ecological data, but on the structural assumptions embedded in network reconstruction.
Despite rapid methodological development in interaction inference, few studies have directly evaluated how alternative reconstruction frameworks condition macroecological conclusions when applied to the same species pool. This gap is particularly consequential for comparative research, where network metrics are routinely interpreted as indicators of environmental filtering, disturbance intensity, evolutionary history, or community stability (Delmas et al. 2018; Allesina and Tang 2012; Poisot et al. 2015). If reconstruction approaches encode distinct structural priors over interaction topology, then differences among communities may reflect reconstruction assumptions rather than ecological processes. Consequently, here we test whether macroecological inference derived from ecological networks is robust to variation in reconstruction framework, asking which aspects of network-based inference are stable across plausible representations of interaction structure and which are intrinsically reconstruction approach dependent.
Deep-time ecosystems provide an especially stringent test of this issue because interactions are not observed directly and must be reconstructed explicitly (Dunne et al. 2008; Dunne et al. 2014; Dunhill et al. 2024; Roopnarine 2006; Karapunar et al. 2026), rendering reconstruction approach assumptions transparent. Against this stringency, here we re-evaluate inferences made by Dunhill et al. (2024) on community structure and extinction dynamics during the Early Toarcian Extinction Event (ETEE; ~183 Ma),a marine biotic crisis in the Early Jurassic, likely caused by a volcanic-driven hyperthermal event (Kemp et al. 2024). Crucially, this re-evaluation allows us to test a pivotal but often overlooked possibility - that the ecological narratives regarding community stability or collapse might be as much a product of the specific reconstruction method chosen as they are of the fossil data itself. By applying alternative reconstruction approaches, we can determine if the Dunhill et al. (2024) conclusions remain robust or if a different choice of reconstruction method would have led to fundamentally different inferences about extinction dynamics. Using four successive assemblages, we reconstruct ecological networks under six contrasting reconstruction classes spanning feasible, realised, and structural representations. For each reconstruction framework, we quantify emergent topology across scales, measure interaction turnover, and simulate disturbance-driven collapse. By holding species composition constant while varying the food web reconstruction approach used, this design isolates the influence of reconstruction constrained structure on inferred food web organisation and extinction dynamics, allowing us to distinguish ecological signals that are robust from those that are reconstruction contingent.
2 Methods
2.1 Study system and fossil data
We used fossil occurrence data from the Cleveland Basin, UK, spanning the upper Pliensbachian to the upper Toarcian. This interval encompasses a major volcanic-driven hyperthermal and marine extinction event. To capture network dynamics across this transition, we defined four successive palaeo-assemblages: pre-extinction (Pliensbachian), post-extinction (Lower Toarcian), early recovery, and late recovery (Middle/Upper Toarcian). Each taxon was characterized using their size and Bambach’s ecospace framework (Bambach et al. 2007), coding for tiering, motility, and feeding mode as per Dunhill et al. (2024). Each assemblage was treated as a community of potentially interacting taxa. The dataset includes 57 taxa across diverse groups (e.g., cephalopods, bivalves, and gastropods).
2.2 Network reconstruction approaches
2.2.1 Conceptual classification of network types
Most palaeontological research (e.g., Fricke et al. (2022); Roopnarine (2006); Shaw et al. (2024)) use reconstruction approaches from within the feasibility space. That is, the reconstructions identify and encode the entire feasible diet of a species to build the network. These methods, however, represent only a subset of the broader spectrum of network construction approaches. Here, we present a suite of methods (Table 1) that enable the construction of a wider range of ecological networks and the exploration of a broader set of ecological questions, provided that their underlying assumptions are compatible with the constraints of the data. Along with one feasibility reconstruction approach we include structural models that create species agnostic networks that are structurally ‘correct’ by assigning links between nodes based on assumptions of link distributions; and realised models that create networks where links between species are constrained based on some form of ‘species choice’ e.g., maximising energy gain.
| Reconstruction Approach | Assumptions | Data needs | Limitation | Network type | Key reference | Usage examples |
|---|---|---|---|---|---|---|
| Random | Links assigned randomly | Species richness, number of links | Parameter assumptions, species agnostic | Structural | Erdős and Rényi (1959) | Null-model comparisons; testing whether observed network structure (connectance, motifs) deviates from random expectations |
| Niche | Species ordered along a ‘niche axis’; interactions interval-constrained | Species richness, connectance | Parameter assumptions, species agnostic | Structural | Williams and Martinez (2008) | Evaluating trophic hierarchy and motif structure; baseline structural predictions |
| Allometric diet breadth model (ADBM) | Energy-maximizing predator diets | Body mass, abundance/biomass | Assumes optimal foraging; does not account for forbidden links | Realised | Petchey et al. (2008) | Predicting realized predator diets; exploring secondary extinctions |
| Allometric trophic network (ATN) | Links constrained by body-size ratios and functional response | Body mass, number of basal species | Assumes only mechanical/energetic constraints | Realised | Brose et al. (2006); Gauzens et al. (2023) | Simulating species loss; evaluating network collapse dynamics |
| Paleo food web inference model (PFIM) | Interactions inferred using trait-based mechanistic rules | Feeding traits | Assumes feeding mechanisms; trait resolution required | Feasibility | Shaw et al. (2024) | Mapping feasible trophic interactions; assessing secondary extinctions |
| Body-size ratio model | Probabilistic assignment of links based on predator–prey size ratios | Body mass | Does not account for forbidden links | Realised | Rohr et al. (2010) | Estimating likely interactions; simulating cascading effects. |
The three body mass-based approaches (ADBM, ATN, Body-size ratio) differ primarily in their underlying ecological assumptions. Although all three approaches use body mass to infer food web structure, they differ in their ecological assumptions. The ADBM is based on energy maximization under optimal foraging theory, the ATN constrains interactions via mechanically optimal consumer-resource size ratios, and the Body-size ratio model defines links probabilistically within a fixed allometric niche. Together, these approaches span bioenergetic, mechanical, and statistical interpretations of size-structured interactions.
2.2.2 Network generation and replication
We evaluated six network reconstruction approaches using the approaches listed in Table 1: Random and Niche models (structural networks); allometric diet breadth (ADBM), allometric trophic network (ATN), and Body-size ratio models (realised networks); and a paleo food web inference model (PFIM; feasibility network). Expanded descriptions of reconstruction approach assumptions, parameterisation, and link-generation rules are provided in Supplementary Material S1. For each community, 100 networks were generated per reconstruction approach per successive community (n = 2400) to capture stochastic variation in link assignment. Where reconstruction approaches required species body mass or trait values, these were sampled within biologically reasonable ranges to preserve relative differences among species. We adopted uniform sampling by default, as alternative distributions (lognormal, truncated lognormal) have negligible impact on topology (Supplementary Material S2; Figure S1). Structural models were parameterised using connectance values drawn from an empirically realistic range (0.07 – 0.34), with species richness held constant. Identical parameter draws were applied across comparable reconstruction approaches within each replicate to ensure comparability. For the Body-size ratio model, we followed the approach of Yeakel et al. (2014) and excluded the latent trait terms.
2.3 Network metrics and structural analyses
We quantified network structure using a suite of macro-, meso-, and micro-scale metrics Table 2, capturing overall network properties, motif structure, and species-level variability. Differences among reconstruction approaches were assessed using a multivariate analysis of variance (MANOVA), with reconstruction approach identity as a fixed factor and the full set of network metrics as response variables. Variance partitioning was further assessed using permutational multivariate analysis of variance (PERMANOVA). Pairwise interaction turnover was quantified using link-based \(\beta\)-diversity was calculated following the framework of Poisot et al. (2012) among the four of the reconstruction frameworks (ADBM, ATN, body-size ratio, and PFIM). We excluded the purely structural Random and Niche models as they are inherently species agnostic. For each pair of reconstructed networks, we represented trophic interactions as binary adjacency matrices and calculated their dissimilarity. Specifically we looked at interaction rewiring among shared species (\(\beta_{OS}\)), which allows separation of differences arising from altered interaction identities among species common to both networks. All calculations were performed for all reconstruction combinations within the same assemblage (time bin).
| Metric | Definition | Scale | Reference |
|---|---|---|---|
| Connectance | \(L/S^2\), where \(S\) is the number of species and \(L\) the number of links | Macro | |
| Maximum trophic level | Prey-weighted trophic level averaged across taxa | Macro | Williams and Martinez (2004) |
| S1 | Number of linear chains, normalised by \(L/S\) | Meso | Milo et al. (2002); Stouffer et al. (2007) |
| S2 | Number of omnivory motifs, normalised by \(L/S\) | Meso | Milo et al. (2002); Stouffer et al. (2007) |
| S4 | Number of apparent competition motifs, normalised by \(L/S\) | Meso | Milo et al. (2002); Stouffer et al. (2007) |
| S5 | Number of direct competition motifs, normalised by \(L/S\) | Meso | Milo et al. (2002); Stouffer et al. (2007) |
| Generality | Standard deviation of normalised generality of all species within network, normalised by \(L/S\) | Micro | Williams and Martinez (2000) |
| Vulnerability | Standard deviation of normalised vulnerability of all species within network, normalised by \(L/S\) | Micro | Williams and Martinez (2000) |
2.4 Extinction simulations and reconstruction approach evaluation
Following Dunhill et al. (2024), we simulated species loss from Pre-extinction networks under trait-based, network-position–based, and random removal scenarios. Species were deleted sequentially, with cascading secondary extinctions allowed to propagate. Simulated post-extinction states were compared to observed (i.e., reconstructed from fossil occurrence data) networks using mean absolute differences (MAD) of food web metrics Table 2 and modified true skill statistics (TSS) calculated separately at the node level (species presence/absence) and link level (presence/absence of interactions between species pairs). Scenarios were ranked within each reconstruction framework based on MAD and TSS performance, and Kendall’s rank correlation coefficient (\(\tau\)) was used to quantify concordance in scenario ordering across reconstruction approaches. Full methodological details are provided in the Supplementary Materials.
2.5 Software and Reproducibility
Ecological network reconstruction and extraction of structural metrics were conducted in Julia v1.11.4 (Bezanson et al. 2017). All statistical analyses, model fitting (MANOVA, PERMANOVA, GAMs), and figure production were performed in R v4.5.2 (R Core Team 2024). The empirical data, derived network datasets and code implementing network reconstruction, extinction simulations, and all analytical workflows are archived at [TODO Zenodo DOI].
3 Results
Results show that reconstruction approaches that appear similar when evaluated using network metrics can yield fundamentally different ecological narratives when interrogated at the level of interactions and extinction dynamics. Across six network reconstruction approaches, inferred food web structure, species interactions, and extinction dynamics differed consistently. Multivariate analyses showed pronounced separation among reconstruction approaches in network metric space. Reconstruction approach explained most of the variance in structural properties, leaving a distinct signature independent of community composition. Notably, approaches that were statistically similar in multivariate structural space often diverged in inferred interactions or extinction dynamics. This demonstrates that structural similarity does not guarantee concordance in species-level diets or trophic roles.
Reconstruction approach substantially influenced inferred extinction dynamics. Temporal trajectories of network collapse, interaction loss, and motif reorganisation differed among approaches. Although node-level extinction ranking were often broadly consistent, link-level outcomes and extinction inferences were highly sensitive to reconstruction assumptions. Together, these results show that ecological inferences drawn from networks depend critically on the reconstruction framework employed.
3.1 Network structure differs among reconstruction approaches
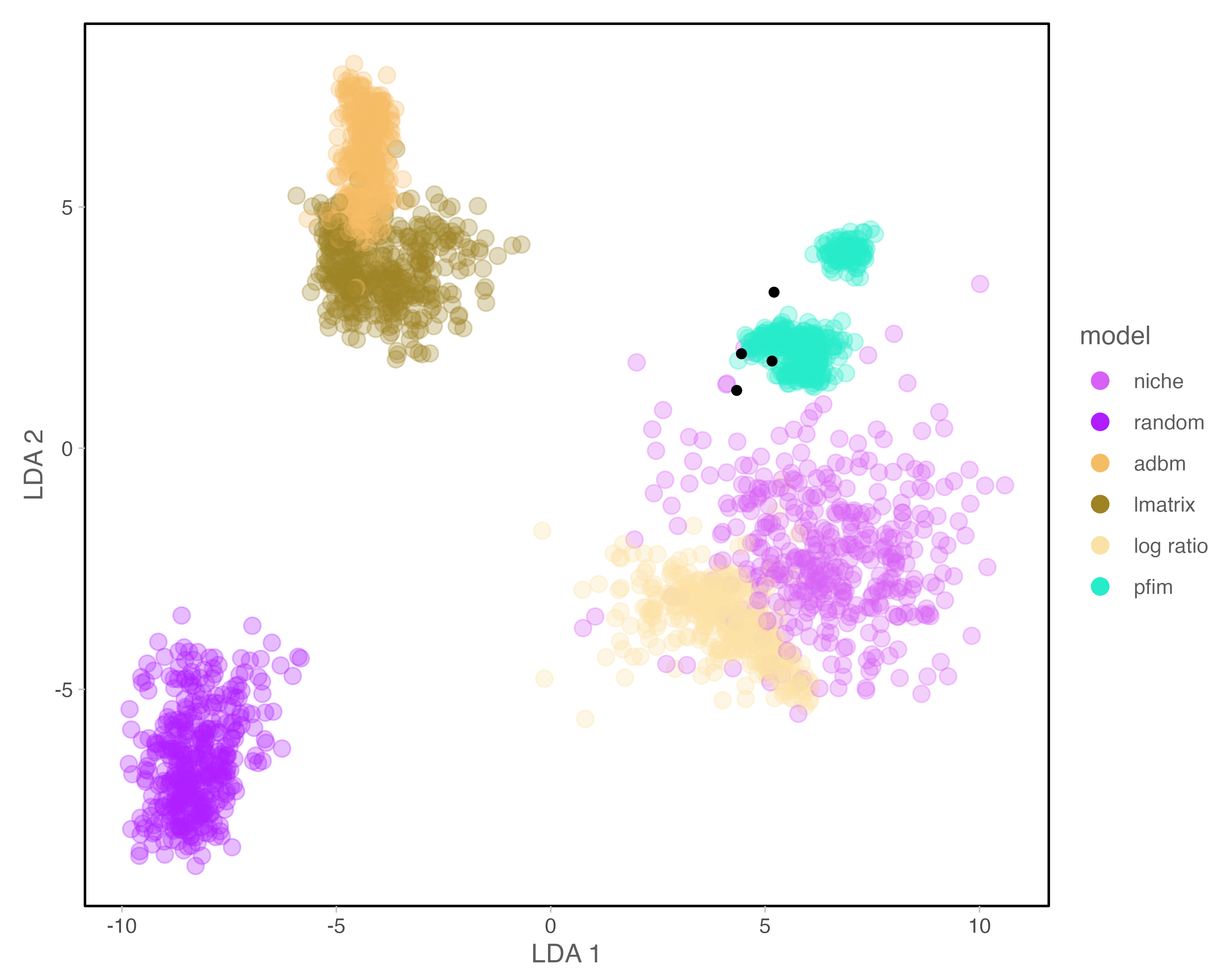
Across six reconstruction approaches, network structure (network properties listed in Table 2) differed significantly (MANOVA, Pillai’s trace = 3.84, approximate \(F_{40,11955}\) = 987.35, p < 0.001), indicating that reconstruction approach systematically alters inferred food web topology. Canonical discriminant analysis identified two dominant axes of variation, explaining 86% of between reconstruction approach variance. LD1 correlated with vulnerability, direct competition motifs, and connectance. LD2 correlated with maximum trophic level and apparent competition motifs, reflecting vertical trophic structure (Figure 1; Table S1, Figure S1). All higher-order canonical variates each explained less than 9% of the remaining variance.
3.1.1 Variance partitioning of network structure
Permutational multivariate analysis of variance revealed that reconstruction framework accounted for the majority of variation in multivariate network structure (\(R^2\) = 0.795, \(p\) < 0.001), whereas temporal turnover across extinction phases explained a comparatively small proportion of variance (\(R^2\) = 0.064, \(p\) < 0.001). The reconstruction approach \(\times\) time interaction contributed a further 7.1% of variance (\(R^2\) = 0.071, \(p\) < 0.001), indicating limited but significant time-dependent divergence among reconstruction frameworks. Thus, differences among reconstruction approaches were more than an order of magnitude greater than structural differences associated with ecological turnover through the extinction sequence, even if the Pliensbachian-Toarcian dataset was characterised with a significant community turnover.
To determine whether the dominance of the reconstruction framework reflected absolute mean shifts among time bins, we repeated the analysis after centring network metrics within each extinction phase. This procedure removes between-phase differences while retaining within-phase structural variation. Even after temporal bin-standardised centring, the reconstruction framework explained 84.8% of multivariate variance (\(R^2\) = 0.848, \(p\) < 0.001). These results demonstrate that the influence of reconstruction approach is not driven by temporal mean differences, but reflects intrinsic divergence among reconstruction frameworks in how ecological interactions are organised.
3.1.2 Statistical Drivers of Network Variation
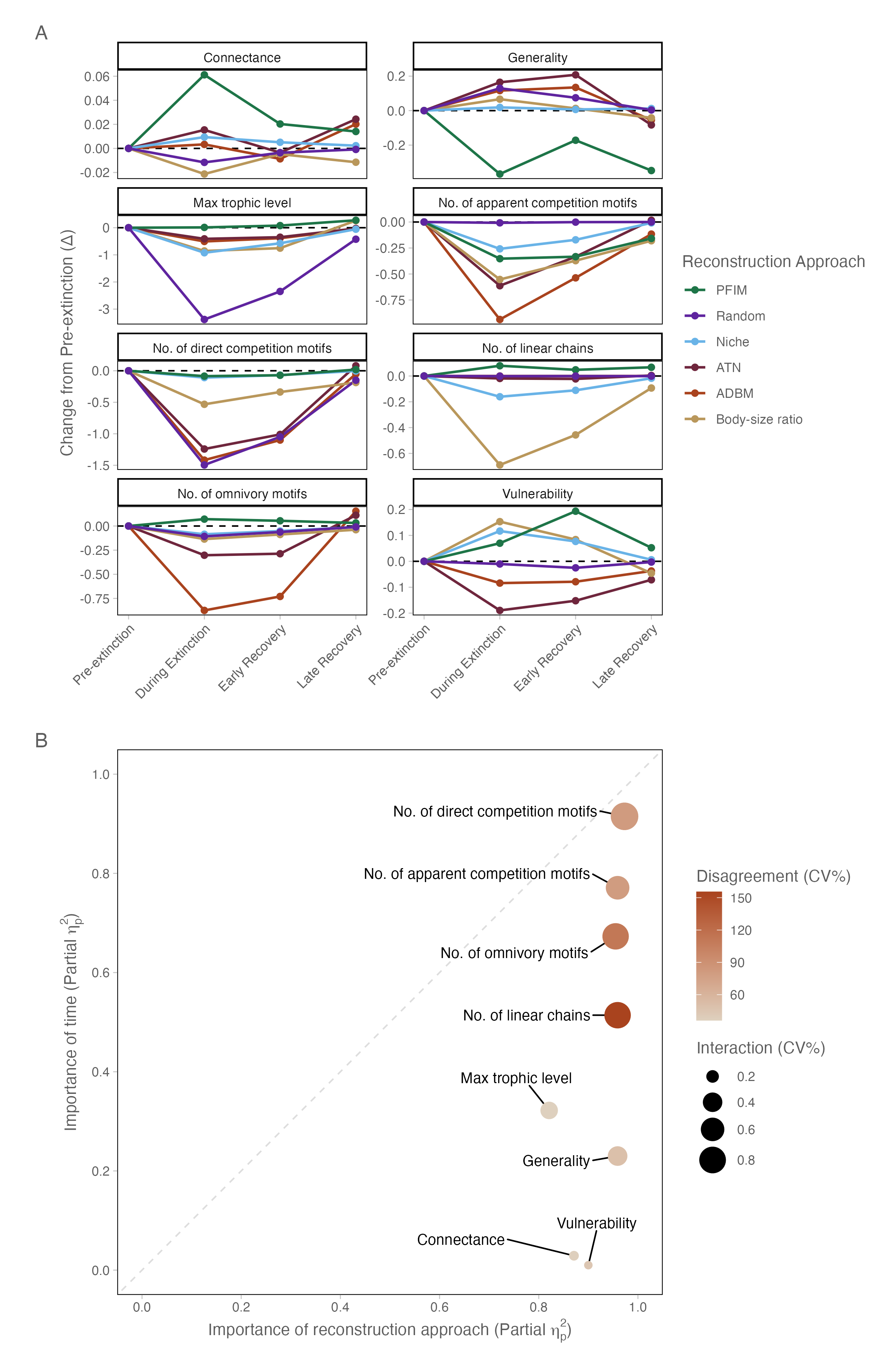
To identify which specific structural properties drive the multivariate separation observed above, we partitioned variance at the level of individual network metrics. Results show that reconstruction choice exerted a significantly stronger influence on network topology than the ecological signal of species loss. In panel A of Figure 2 it is clear that for certain network metrics reconstruction approaches predict different responses across time e.g., connectance, as well as differing magnitudes of change e.g., the number of apparent cmpetition motifs. A two-way factorial ANOVA across all eight network metrics confirmed that the reconstruction approach was the dominant driver of variance, with partial eta-squared values (\(\eta^{2}_{p}\)) consistently exceeding 0.82 and reaching 0.97 for meso-scale motifs (Figure 2, panel B; Table S3). While the extinction event (time bin) significantly altered network structure (p<0.001), its relative importance remained secondary, typically explaining a smaller fraction of the total topological variation. This is clear in panel B of Figure 2 where all metrics are within the bottom-right (reconstruction approach-dominated) quadrant of this space, emphasising that framework assumptions outweigh the ecological signal of species loss. Furthermore, the high inter-reconstruction approach Coefficient of Variation (CV) observed for some metrics (Table S4, Figure S4) highlights a sensitivity. The properties that are influenced by time are also those upon which the reconstruction approaches disagree most profoundly. Demonstrating that our understanding of structural food web collapse in the fossil record is highly contingent on the chosen reconstruction framework, particularly when examining complex trophic pathways beyond simple macro-scale properties like connectance.
3.1.3 Inferred pairwise interactions vary widely among reconstruction approaches
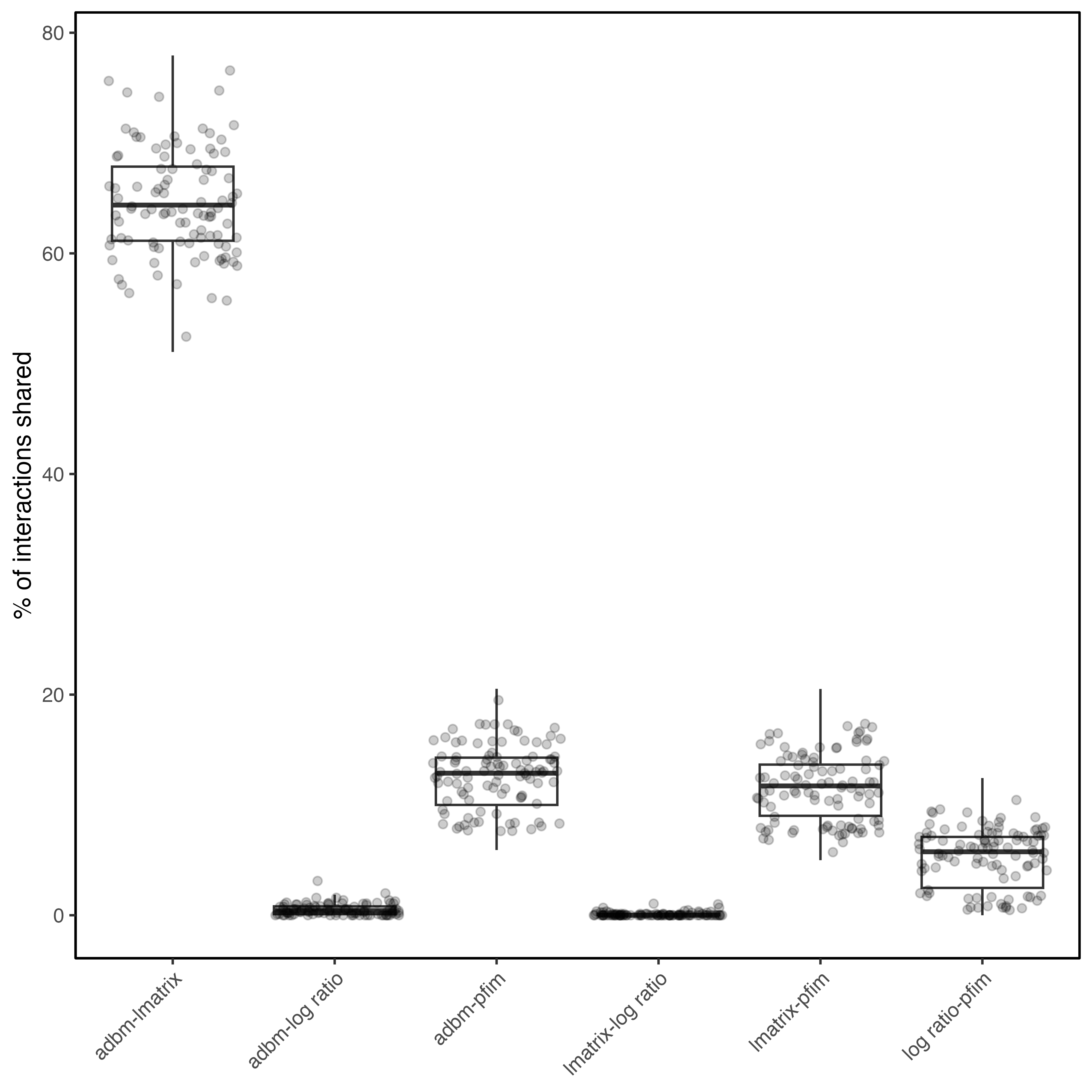
Despite some network reconstruction approaches showing similar netwrok metrics, specific pairwise interactions often differed. Pairwise \(β_{OS}\) revealed that certain reconstruction approach pairs shared very few links Figure 3. Size-based models (ADBM, ATN) were broadly similar due to shared sole reliance on body-size constraints, whereas the Body-size ratio model exhibited consistently higher differences to other reconstruction approaches. PFIM showed intermediate overlap with body mass-based models. These results demonstrate that agreement in overall network structure does not guarantee concordance in species-level interactions.
3.2 Reconstruction approach choice influences inferred extinction dynamics
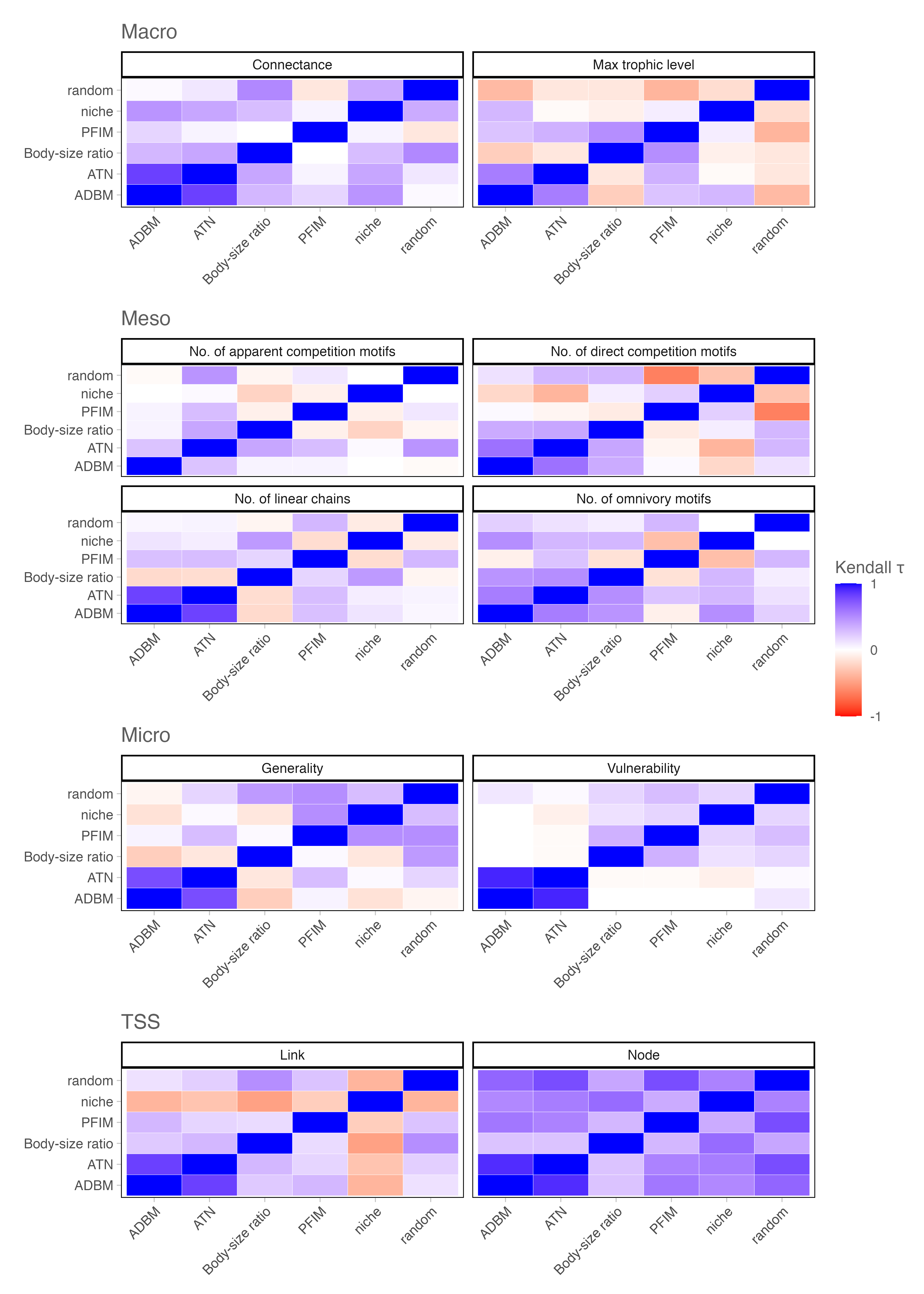
To evaluate how reconstruction approach choice affects inferred extinction dynamics, we compared simulated post-extinction networks to observed networks using mean absolute differences (MAD) for network-level metrics and true skills statistics (TSS) for node- and link-level outcomes Figure 4. Across reconstruction approaches, MAD-based rankings were generally positively correlated (Kendall’s \(\tau\) ≈ 0.13 across structural metrics), indicating weak but generally positive concordance on the relative importance of extinction drivers despite substantial differences in reconstructed network structure. However, agreement within the allometric models differed from patterns observed for reconstructed network structure.
Node-level TSS rankings were similarly consistent across network reconstruction approaches (Kendall’s \(\tau\) = 0.26 - 0.90), reflecting broadly comparable node-level removal sequences. In contrast, link-level outcomes were far more variable (Kendall’s \(\tau\) = −0.48 - 0.29), highlighting that inferences about which interactions are lost, retained, or re-established during collapse and recovery are highly reconstruction approach contingent. Together, these results suggest that while alternative reconstruction approaches converge on similar species-level extinction patterns, the inferred pathways of interaction loss and cascading dynamics depend strongly on both reconstruction approaches.
4 Discussion
4.1 Network reconstruction is not neutral: structural priors shape ecological theory
Food web ecology has long treated network reconstruction as a technical step preceding ecological analysis. Once a network is assembled (whether from observation, inference, or simulation) its properties are typically analysed as reflections of underlying ecological organisation. Implicit in this workflow is a powerful assumption - that reconstructed networks provide structurally comparable representations of ecological communities, such that differences in connectance, trophic structure, motif composition, or robustness primarily reflect biological variation.
This assumption is particularly critical to evaluate within the context of deep-time palaeoecological data. Because interactions in fossil ecosystems are never observed directly, they must explicitly be formed through some form of reconstruction approach. This necessity renders the underlying assumptions transparent but also makes the resulting ecological narratives highly susceptible to the constraints inherent in the chosen reconstruction framework. In these settings the risk is not just incomplete data, but the potential for methodological artefacts to be misinterpreted as genuine macroevolutionary or palaeoecological signals. Consequently, deep-time studies offer a unique and stringent testing ground for determining whether community-level responses (such as stability or collapse during mass extinctions) are robust features of the ecosystem or merely byproducts of how we choose to construct the links between species.
Reconstruction framework explained the overwhelming majority of variance in inferred food web topology, far outweighing the influence of temporal turnover across extinction and recovery phases. Across an identical regional taxon pool, alternative reconstruction approaches generated distinct structural signatures that occupied non-overlapping regions of multivariate space Figure 1, demonstrating that divergence among reconstruction approaches reflects intrinsic differences in how interactions are organised rather than temporal shifts in community composition. Even after centring metrics within time bins (extinction phases) to remove between-bin mean differences, reconstruction approach remained the dominant driver of structural variation. These results indicate that reconstruction approaches impose distinct ‘structural priors’ on ecological inference. These priors are not subtle; they propagate into emergent topology, species roles, and predictions of disturbance dynamics. Network structure is therefore not solely a property of ecological communities, but jointly determined by ecological data, reconstruction assumptions, and level of organisation (Strydom et al. 2026; Gauzens et al. 2025).
Crucially, this dominance was not confined to multivariate summaries. Variance partitioning at the level of individual network properties revealed that reconstruction approach overwhelmingly structured specific ecological metrics, with partial η² values exceeding 0.9 for meso-scale motif composition and remaining consistently high across macro- and micro-scale properties. Thus the imprint of reconstruction framework is visible not only in aggregate topology but in the very structural features often interpreted as ecological signals — trophic height, competition, and variability in species roles. Notably, the few properties that exhibited detectable temporal sensitivity were also those with the greatest inter-reconstruction approach disagreement, indicating that temporal trends is most difficult to disentangle when reconstruction frameworks diverge most strongly. These results suggest that structural assumptions do not merely shift networks within a shared architectural space; they condition the specific patterns through which ecological change is perceived and interpreted.
This has direct implications for the interpretation of comparative network studies. Feasible, realised, and structural reconstruction approaches encode different assumptions about constraint, optimisation, and topology, with these assumptions propagating into emergent metrics and dynamical predictions (Allesina and Tang 2012; Curtsdotter et al. 2011; Dunne et al. 2002; Poisot and Gravel 2014; Michalska-Smith and Allesina 2019). When networks reconstructed under different classes are compared across spatial gradients, disturbance regimes, or evolutionary transitions, part of the observed variation may derive from reconstruction choice rather than ecological process. Without explicit standardisation or sensitivity analysis, methodological heterogeneity can be mistaken for biological signal. Food web ecology has devoted substantial effort to understanding how topology shapes dynamics; comparatively less attention has been paid to how reconstruction method shapes topology. Our findings indicate that these two questions cannot be separated.
4.2 Scale-dependent robustness in network-based inference
Importantly, reconstruction sensitivity was not uniform across network scales (macro-, mesio-, micro- level properties). Node-level extinction rankings were broadly consistent among reconstruction approaches, whereas interaction-level outcomes and cascade trajectories were highly contingent on reconstruction approach. The predominance of reconstruction framework over temporal turnover (~80% vs. 6% variance explained) illustrates why coarse-grained patterns like node-level extinction rankings are more robust. reconstruction approach-imposed structure dominates the overall topology, leaving interaction dynamics highly contingent on framework choice. This asymmetry reveals a context-dependent pattern of robustness. Coarse-grained macroecological patterns (such as the vulnerability of a community to collapse) can emerge from multiple plausible interaction architectures. By contrast, fine-grained inferences about which links are lost, retained, or reorganised depend strongly on how interactions are inferred.
This distinction challenges a central ambition of food web ecology: the use of detailed interaction structure to diagnose mechanisms of stability and collapse. Our findings suggest that while coarse-grained patterns might be shared across methods, fine-grained mechanistic narratives (such as the specific pathways of interaction loss) are much more precarious. This implies that had Dunhill et al. (2024) selected a different reconstruction method, the resulting inferences regarding the drivers of extinctions and subsequent recovery could have pointed to entirely different ecological mechanisms. If interaction-level cascade pathways vary substantially across equally plausible reconstructions, then mechanistic narratives derived from a single inferred topology may overstate their precision (Dunne et al. 2002; Allesina and Tang 2012; Curtsdotter et al. 2011). The apparent determinism of extinction cascades may therefore partly reflect reconstruction-imposed structure rather than ecological inevitability.
For macroecology, this metric dependence clarifies where network-based inference is accurate. Aggregate properties may be comparatively robust to reconstruction assumptions, whereas conclusions about interaction turnover, motif reorganisation, or fine-scale trophic dynamics are intrinsically uncertain. Recognising this asymmetry is essential if network analyses are to inform comparative synthesis across space and time.
Taken together, these results underscore that network reconstruction is not a neutral preprocessing step but an additional part of the hypothesis-generating process in which each reconstruction approach encodes a distinct set of ecological assumptions. The inferred topology and dynamics of a food web therefore reflect not only ecological data, but the theoretical assumptions embedded in the reconstruction framework. Disagreement among reconstruction approaches does not imply that any single approach is ‘wrong’, but rather that different reconstruction approaches capture different facets of ecological reality (Stouffer 2019; Petchey et al. 2011). Disagreement among reconstruction approaches does not imply that any single approach is ‘incorrect’. Rather, different reconstruction approaches capture different facets of ecological constraint—trait compatibility, energetic optimisation, or topological regularity. The critical point is that these facets are not interchangeable.
This perspective reframes reconstruction choice as part of hypothesis specification. Researchers must align reconstruction approaches with the ecological signals of interest (whether potential interactions, realised diets, or macro-scale structural properties) rather than treating network reconstruction as a technical convenience. Viewed through the lens of accuracy and precision, our results indicate that some network-based inferences are relatively robust across reconstruction approaches, whereas others remain intrinsically uncertain. High-level extinction rankings were broadly convergent, suggesting relative accuracy at coarse resolution, but interaction-level details and temporal cascade dynamics diverged substantially, indicating limited precision in reconstructing the fine structure of collapse. Recognising and explicitly accounting for this distinction is essential if food web ecology is to move beyond descriptive reconstruction toward rigorous comparative inference.
4.3 Implications for comparative biogeography and global change research
Network approaches are increasingly applied to examine how ecological organisation varies across latitudinal gradients, environmental filters, disturbance regimes, and climate-driven transitions (Gilman et al. 2010; Tylianakis et al. 2008). In global change ecology, networks are used to project vulnerability under warming, quantify rewiring of interactions, and assess stability under species loss (e.g., Hao et al. 2025; Marjakangas et al. 2025). These studies frequently interpret variation in connectance, trophic height, interaction \(\beta\)-diversity, or robustness as indicators of ecological differentiation among regions or time intervals (e.g., Pellissier et al. 2018; Trøjelsgaard and Olesen 2016). Our results show that such differences can systematically alter inferred topology and disturbance dynamics even when species composition is held constant. This suggests that apparent differences in network structure across spatial or climate gradients may reflect variation in reconstruction as much as ecological process.
Deep-time palaeo food webs provide a complementary perspective because they capture ecosystem responses to large-scale environmental perturbations and extinction events under past climate change (e.g., Dunhill et al. (2024); Karapunar et al. (2026); Smith et al. (2025)). Fossil networks therefore represent natural experiments for evaluating resilience, trophic reorganisation, and recovery following extreme environmental change. Studies of palaeo food webs have demonstrated how network structure mediates extinction cascades and post-disturbance reassembly (Roopnarine 2006; Dunne et al. 2008), providing empirical constraints on long-term ecological stability.
However, our results emphasise that even in deep-time systems structural conclusions remain sensitive to reconstruction approaches. Treating reconstructed networks as ensembles rather than single deterministic representations provides a more transparent framework for incorporating uncertainty into comparative macroecology and for using palaeo data to inform expectations about modern climate-driven reorganisation.
4.4 Toward a more explicit modelling paradigm in food web ecology
The broader implication is not that any single reconstruction framework is ‘correct’ or ‘incorrect’. Rather, each reconstruction approach represents a distinct hypothesis about how interactions are constrained—by trait compatibility, energetic optimisation, or topological regularity (Petchey et al. 2011). Food web reconstruction is therefore theory-laden. Making this explicit shifts reconstruction from a preparatory step to a central component of ecological modelling.
A mature modelling paradigm in food web ecology would treat reconstruction approaches as testable, incorporate probabilistic link inference where possible, and quantify the sensitivity of macroecological conclusions to alternative representations of interaction structure. Such an approach aligns with recent advances in probabilistic and ensemble network modelling and would strengthen the interpretability of network-based inference under global change (Banville et al. 2025; Poisot et al. 2016).
5 Conclusions
Ecological network reconstruction is not a neutral technical procedure but a theoretical act that shapes ecological inference. By applying six contrasting reconstruction frameworks to an identical species pool, we show that different reconstruction approaches systematically influence inferred food web topology, interaction identity, and disturbance dynamics. Some coarse-grained patterns, such as relative species vulnerability, are comparatively robust across representations. In contrast, fine-scale interaction structure and cascade pathways are highly contingent on reconstruction approach. The reliability of network-based inference is therefore scale dependent.
These results challenge the implicit assumption that reconstructed networks are comparable across systems — whether comparing modern communities across environmental gradients or fossil assemblages across extinction intervals. When reconstruction frameworks differ, variation in connectance, trophic organisation, robustness, or interaction turnover may reflect embedded reconstruction approaches as much as ecological processes. Network reconstruction should thus be treated as an explicit component of hypothesis specification in comparative macroecology and biogeography.
No single reconstruction approach captures the full complexity of ecological organisation, but neither are alternative reconstruction approaches interchangeable. Aligning reconstruction framework with inferential goals, standardising approaches across comparative studies, and incorporating ensemble or probabilistic representations will be essential for strengthening the interpretability of network analyses across spatial and temporal gradients, including efforts to use deep-time systems to inform expectations under contemporary climate change. As ecological networks increasingly inform global change research, recognising network reconstruction as fundamental determinants of inference is critical for advancing food web ecology from descriptive reconstruction toward rigorous comparative synthesis.
References
Citation
@online{strydom2026,
author = {Strydom, Tanya and Karapunar, Baran and A. Dunne, Jennifer
and Hull, Pincelli and T.S. Little, Crispin and Pimiento, Catalina
and Ridgwell, Andy and P. Beckerman, Andrew and Dunhill, Alexander},
title = {Network Reconstruction Approach Conditions Ecological
Inference in Food Web Reconstruction},
date = {2026-04-01},
langid = {en},
abstract = {Aim Ecological networks are widely used to compare
community structure, stability, and responses to disturbance across
environmental gradients. However, many networks (particularly those
assembled from incomplete interaction data) require model-based
reconstruction. We test how alternative reconstruction frameworks
condition ecological inference by quantifying their effects on
network structure and disturbance dynamics. Location Cleveland
Basin, United Kingdom. Time period Toarcian extinction event (Early
Jurassic, late Pliensbachian–late Toarcian, \textasciitilde183 Ma).
Major taxa studied Shelly marine macrofossils. Methods We
reconstructed four successive assemblages from an identical species
pool using six contrasting food web reconstruction approaches
spanning feasible (trait-based), realised (allometric and
energetic), and structural (topological) network representations.
For each community and reconstruction approach, 100 replicate
networks were generated. We quantified macro-, meso-, and
micro-scale network properties and assessed differences among
reconstruction approaches using multivariate analyses. Pairwise
interaction turnover was measured using link-based beta diversity.
We then simulated species loss under multiple disturbance scenarios,
allowing cascading extinctions, and compared predicted community
states using mean absolute differences and rank concordance metrics
between reconstruction approaches. Results Reconstruction framework
strongly influenced inferred network topology, generating distinct
structural signatures independent of species composition.
Reconstruction approaches that were similar in network metrics often
diverged in species-level interactions, with high β-turnover among
inferred link sets. During extinction simulations, scenario rankings
were broadly consistent at the network level, but interaction-level
outcomes and cascade dynamics varied substantially. Main conclusions
Network reconstruction functions as a structural prior that
conditions ecological inference. While some aggregate patterns are
robust across reconstruction approaches, detailed interaction-level
dynamics are highly contingent on reconstruction approach.
Comparative network studies across spatial or environmental
gradients should therefore align reconstruction framework with
inferential goals and explicitly evaluate sensitivity to
reconstruction assumptions.}
}